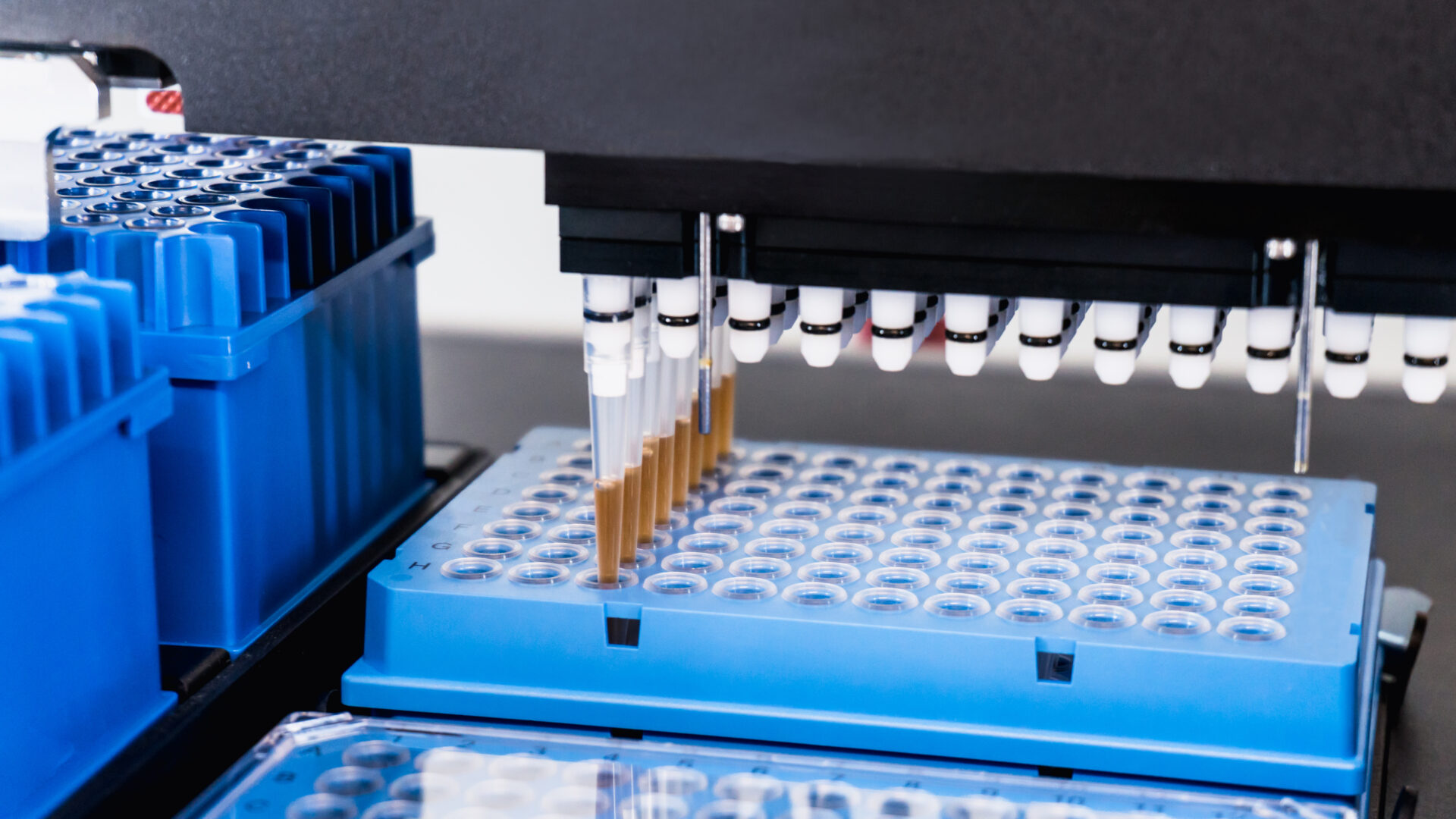
Next-Gen Sequencing is an important diagnostic tool for veterinarians.
DNA (also known as deoxyribonucleic acid) has long been nicknamed the “blueprint for life”, providing the genetic codes that make existence for all organisms possible [1]. From animals to bacteria to fungi, each organism’s genetic code is what makes them unique and can also provide an abundance of information for clinical uses. Specifically, the development of high throughput molecular technologies alongside bioinformatics analyses in the 21st century has radically changed the abilities of veterinarians to give their patients more informed diagnoses and treatments [2]. NGS is a “high-resolution tool that provides veterinary diagnostic laboratories with the ability to undertake swift and flexible responses to emerging infectious diseases and unexpected pathogen variants” [2].
If you suspect your pet is suffering from an infection, it is recommended that you make an appointment with your veterinarian to diagnose and provide a treatment plan tailored to your pet’s needs as soon as possible!
What can DNA tell veterinarians about your pet’s infection?
DNA is composed of two parallel strands made from a phosphate sugar backbone and is connected by chemical bonds between bases (adenine bonds to thymine, and cytosine bonds to guanine), which creates a double helix [1]. NGS has allowed for the characterization of complete microbial communities, even those without prior knowledge [3]! The analysis of metagenomics data can not only help identify and quantify the microbiome of interest, but can also be function based and identify coding gene diversity [2]. This means that NGS diagnostics can help screen for virulence associated, antibiotic resistance genes, and vitamin production-associated genes in the microbial communities [2]!
So, how does it work?
For NGS diagnostics, microbial, genomic DNA or RNA (converted to cDNA) is extracted from clinical samples, such as urine, feces, blood, or skin/ear/nose/throat swabs, then purified and tagged. The resulting DNA molecules are copied thousands of times and massively sequenced in parallel. Thus, millions of DNA sequence reads from all microbes in a sample are able to be examined at once. These DNA reads are then trimmed, host DNA is filtered out using special software, overlapping reads are assembled into long sequences and the genes are compared to genomic databases to assign the microbial species identity and associated pathogenicity and predict antimicrobial activity based on resistance genes. This process provides a complete picture of which microbes are present and allows the quantification of each microbial species at the time and sight of sample collection from the patient. NGS-based diagnostic tools are increasingly used in veterinary practices to further improve infection diagnostics, while also aiding good antibiotic stewardship by selecting the most useful antibiotics. These types of tests are offered by specialty labs, like the MiDOG labs pet microbiome test.
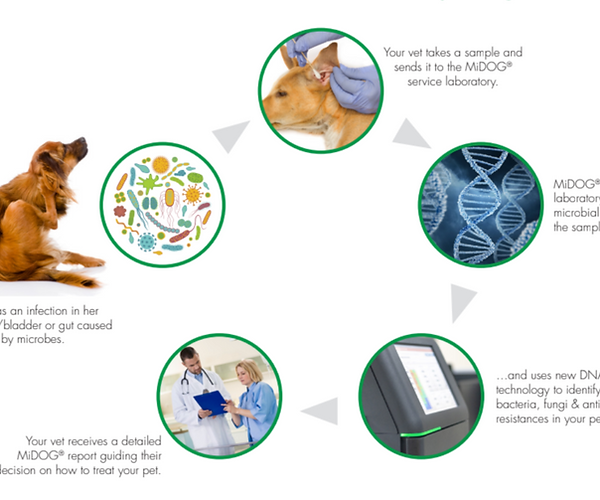
The image above depicts the MiDOG All-in-One Microbial Test workflow.
Why is Next-Gen Sequencing Better Than Cultures or PCR?
Historically, culture-based methods and PCR techniques have been used to assess potential emerging infections in veterinary medicine, but there are notable diagnostic shortcomings for both. Only one percent of all microbes can actually be cultured using standard culturing methods, which lack the sensitivity and ability to cultivate difficult to grow pathogens [4]. While PCR provides more rapid results, the diagnostic value of these tests is limited because they are only able to target a known set of pathogens.
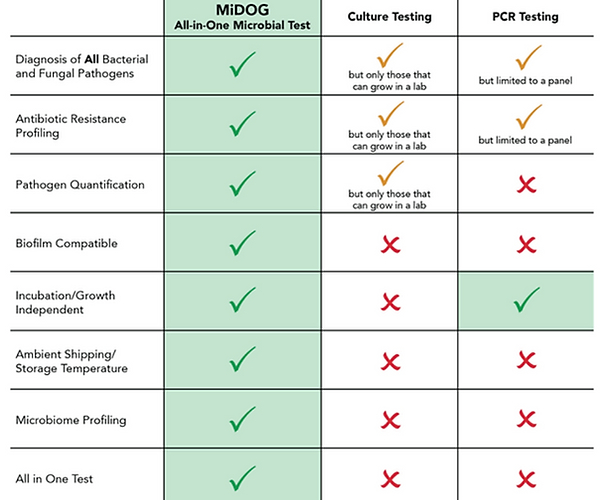
The MiDOG All-in-One Microbial test is superior to culture and PCR testing.
Next-Gen Sequencing has increasingly helped researchers and veterinarians characterize the skin, oral, ear, gut, and fecal microbiome of pets. This translates to direct treatment. For example, MiDOG NGS technology helped Nola, a 3-year-old Affenpinscher that was suffering from a urinary tract infection. Her veterinarian, Dr. Michael Morgan from Tustin, CA, collected her urine sample via cystocentesis and sent it out for both culture testing and to MiDOG for NGS testing. The culture results were a “no growth”, meaning the amount of bacteria growth fell below the threshold for a urinary tract infection. However, the MiDOG All-in-One Test showed a huge overgrowth of Proteus mirabilis, which is a facultatively anaerobic bacteria that the culture test missed. With this informed knowledge, Dr. Morgan was able to come up with the best treatment plan for Nola and help her recover!
The MiDOG All-in-One Microbial Test offers comprehensive diagnostic information for all different types of animals suffering from an infection. Utilizing NGS technology to detect and quantify all microbial DNA through untargeted and comprehensive sequencing and quantitative comparisons to reference databases, the MiDOG NGS technology provides a useful opportunity to shed light on the microbial makeup of your pet’s microbiome for clinical application. The MiDOG microbiome test is a microbial identification test grounded on scientific research that provides veterinarians DNA evidence for guided treatment with proven results.

Find out if your vet uses MiDOG before you book your next appointment!
References:
- Weir, M., Ruotsalo, K., & Tant, M. (2022). Dna Testing | VCA Animal Hospitals. Retrieved 6 May 2022, from https://vcahospitals.com/know-your-pet/dna-testing
- Van Borm, S., Belák, S., Freimanis, G., Fusaro, A., Granberg, F., & Höper, D. et al. (2014). Next-Generation Sequencing in Veterinary Medicine: How Can the Massive Amount of Information Arising from High-Throughput Technologies Improve Diagnosis, Control, and Management of Infectious Diseases?. Veterinary Infection Biology: Molecular Diagnostics And High-Throughput Strategies, 415-436. doi: 10.1007/978-1-4939-2004-4_30
- Behjati, S., & Tarpey, P. (2013). What is next generation sequencing?. Archives Of Disease In Childhood – Education &Amp; Practice Edition, 98(6), 236-238. doi: 10.1136/archdischild-2013-304340
- Krumbeck, J., Holden, N., & Malka, S. (2022). The Importance of Next-Generation Sequencing in Avian Veterinary Medicine – LafeberVet. Retrieved 6 May 2022, from https://lafeber.com/vet/the-importance-of-next-generation-sequencing-in-avian-veterinary-medicine/
Categories: Next-Gen DNA Sequencing Technology, Veterinarian Guides, Veterinary Best Practices, Veterinary Dermatology